R and RStudio
Andreas Handel
2023-06-14 15:14:51.680529
Overview
This is a very quick introduction to R and RStudio, to get you set up and running. We won’t go deep into coding, but we need R to use the DSAIRM package. R is a very useful tool to be familiar with if you want to get into modeling, or if you want to do data analysis. As such, having a bit of familiarity is good.
Learning Objectives
- Know some of the strengths and weaknesses of R.
- Have R and Rstudio up and running on your computer.
- Know how to install and load R packages.
About R
Like every programming language, R has its advantages and disadvantages. Feel free to do a web search on that topic, and you will encounter tons of people with tons of opinions. Some of the features that are useful to us are:
- R is Open Source, FREE and cross-platform.
- R is a “high-level” programming language, relatively easy to learn (compared to Fortran, C, etc.).
- R comes with many integrated functions.
- R is great for statistics, data analysis, and graphics.
- R is a full programming language and can do a lot of things, including running infectious disease models.
- The R Community is very dynamic, helpful and welcoming.
- Through R packages, it is easy to get lots of state-of-the-art algorithms.
- Documentation and help files for R are generally good.
While we use R, it is not the only option for building and exploring infectious disease models. Maybe the most similar to R, and widely used, is Python, which is also free. There is also commercial software that can be used (e.g., Matlab, Mathematica). Other more general programming languages are suitable too (e.g., C, Fortran, Java, Julia). Depending on your future needs or jobs, you might have to learn one or several of those languages. In general, knowing some coding is a very useful skill, and I strongly encourage you to learn some programming. The good news is that while all programming languages are somewhat different, they all share general ways of thinking and structuring code. So once you understand a specific concept (e.g., variables, loops, branching statements or functions), it applies to all those languages. Thus, learning a new programming language is much easier once you already know one. R is a good one to get started with.
Installing R and RStudio
- If you haven’t already, install R first. You can pick any mirror you like. If you already have R installed, make sure it is a fairly recent version. Currently, DSAIRM requires R 4.0 or newer. If yours is older, you need to update (install a new R version).
- Once you have R installed, install the free version of RStudio Desktop. Again, make sure it’s a recent version.
Installing R and RStudio should be fairly straightforward. Everything should work on all the standard operating systems (Windows, Mac, and even Linux). If you want detailed instructions this reading provides more detailed steps. If things don’t work, ask for help on Slack.
R packages
Most of the functionality and features in R come in the form of
add-on packages. There are tens of thousands of packages available, some
big, some small, some well documented, some not. We’ll be focusing on
the DSAIRM package in this course. Of course, you are free
to install and use any package you come across and want to try.
Installing R packages from CRAN
The main official place for R packages is the CRAN
website. If you are interested in packages on a specific topic, the
CRAN task views
provide curated descriptions of packages sorted by topic.
To install a package from CRAN, go to the R prompt at the bottom left
of your RStudio session and type
install.packages("PACKAGENAME"). The figure shows an
example where I installed a package called learnr. Often, a
package needs other packages to work (called dependencies), and they are
installed automatically. It usually doesn’t matter if you use a single
or double quotation mark around the name of the package. Note
that R cares about capitalization, so you need to get the upper and
lower case exactly right. Otherwise, it won’t work.
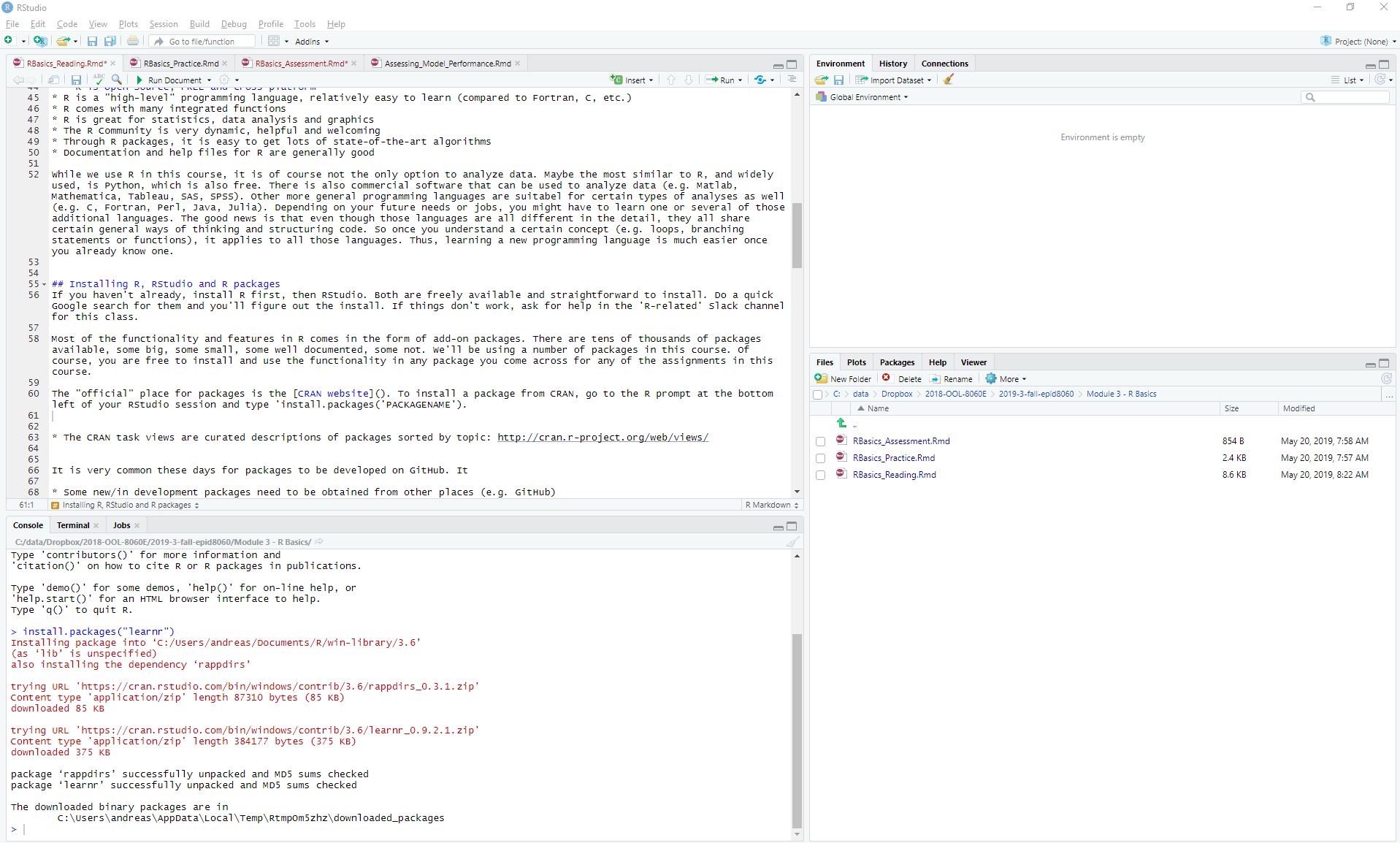
Installing an R package. Right-click on the image to enlarge it.
Try installing a package yourself. Open RStudio. Then go to the R
prompt (the > symbol) in the lower-left corner and
type
install.packages('shiny')In RStudio, you can also install (and update/remove) packages by clicking on the ‘Packages’ tab in the bottom right window.
Installing R packages from GitHub
It is very common these days for packages to be developed on GitHub.
It is possible to install packages from Github directly. Those usually
contain the latest version of the package, with features that might not
be available yet on the CRAN website. Sometimes, in early development
stages, a package is only on Github until the developer(s) feel it’s
good enough for CRAN submission. So installing from Github gives you the
latest. The downside is that packages under development can often be
buggy and not working right. To install packages from Github, you need
to install the remotes package and then use the
install_github function. Here is an example, installing Hadley Wickham’s emo package,
which allows you to easily insert emojis into RMarkdown files and which
is - as of this writing - not available from CRAN.
install.packages('remotes') #needed to install from GitHub
library('remotes')
install_github('hadley/emo')Loading packages
You only need to install a package once (from CRAN or GitHub or some
other way), unless you upgrade/re-install R. Once installed, you still
need to load the package before you can use it. That has to happen every
time you start a new R session. You do that using the
library() command (an alternative is require()
but I recommend library()). For instance to load the
DSAIRM package, type
library('DSAIRM')Some packages show messages when you load them, and others don’t. In
this case, the package tells you that you can start using it by typing
dsairmmenu() into the R console. You can do that now, or go
straight to the next reading, which will talk about DSAIRM in more
detail.
Learning RStudio
While one can use R and do pretty much every task, including all the ones we cover in this course, without using RStudio, RStudio is very useful, has lots of features that make your R coding life easier and has become pretty much the default integrated development environment (IDE) for R. Since RStudio has lots of features, it takes time to learn them. You don’t really need to know much about RStudio for this course. But if you are interested, a good resource to learn more about RStudio are the RStudio Essentials collection of videos.